Virtual lungs to understand the dynamics of tuberculosis lesions within the lungs
The study focusses on understanding the mechanisms that keep the infection latent without the infected person developing the disease. 90% of people infected never develop the active disease, understanding this mechanism so that we can identify them is key to eradicating tuberculosis. The research has been carried out in virtual lungs, developed by computational modelling techniques. These are the first results to be published by the 3Rs Programme at the Centre for Comparative Medicine and Bioimage (CMCiB) which aims to minimize the use of animals in pre-clinical research.
The researchers of the Experimental Tuberculosis Unit (UTE) of the IGTP, led by Pere-Joan Cardona, have published the first results of a computational model that aims to reproduce the dynamics of tuberculosis lesions in a virtual lung. The model has been developed within the 3Rs Programme of the Centre for Comparative Medicine and Bioimage (CMCiB) of the IGTP, supported by the "la Caixa" Foundation, with Clara Prats (Researcher at the UPC Universitat Politècnica de Catalunya BarcelonaTech (UPC) and CMCiB) and Martí Català of the CMCiB, first and last authors on the paper respectively. The results are published in the journal Plos Computactional Biology.
Today tuberculosis (TB) is still one of the ten principal causes of death in the world, in 2017 1,6 million people died because of the disease. We know that tuberculosis is caused by Mycobacterium tuberculosis, a bacterium that infects the air sacs, or alveoli, of the lungs. However, we also know that 90% of people will never develop the active disease. The fact that we do not know the main factors that trigger the disease in the remaining 10% of cases is one of the main obstacles to its eradication. This study aims to understand the mechanisms that allow the disease to remain latent, especially those related to endogenous reinfection.
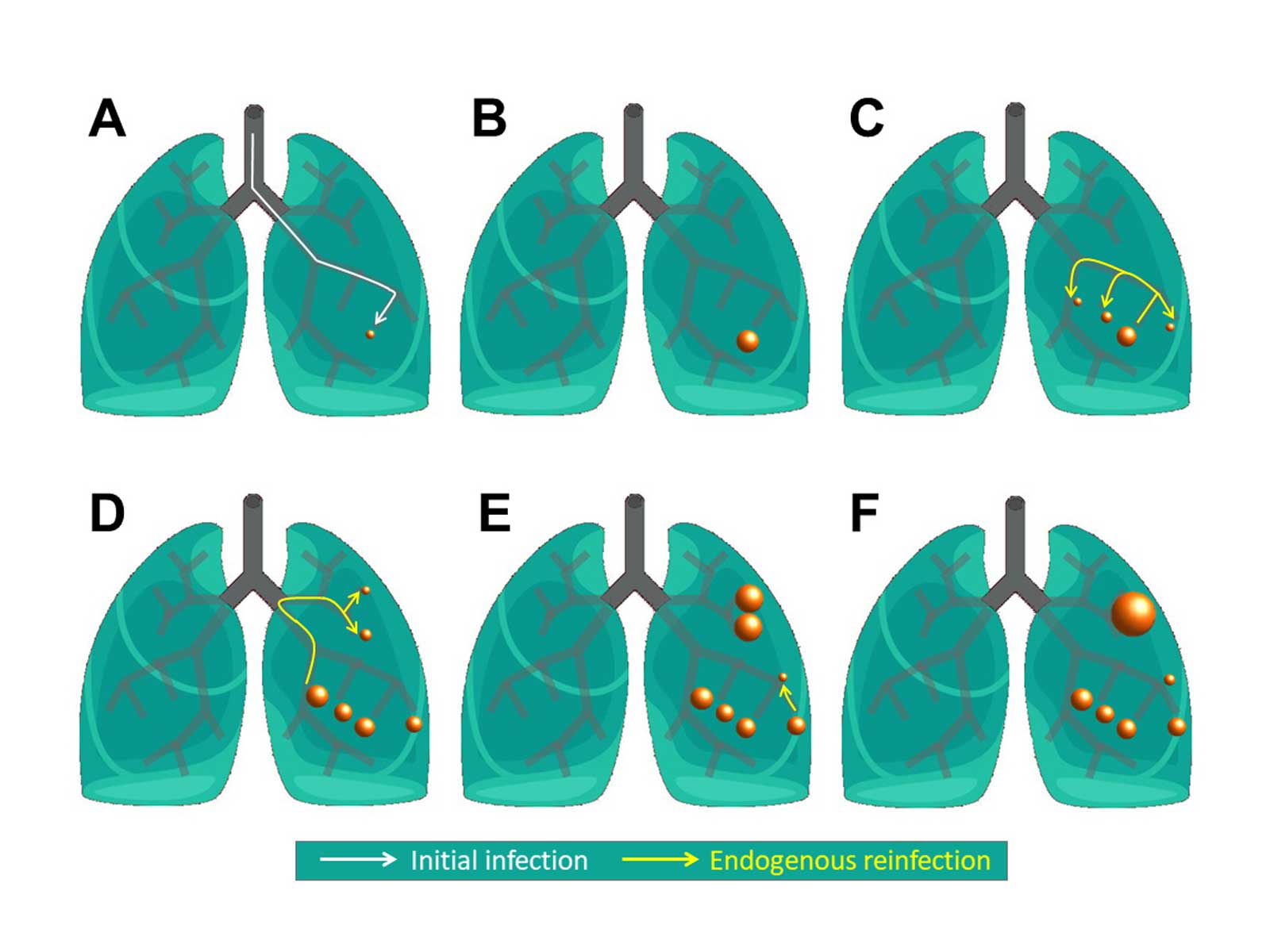
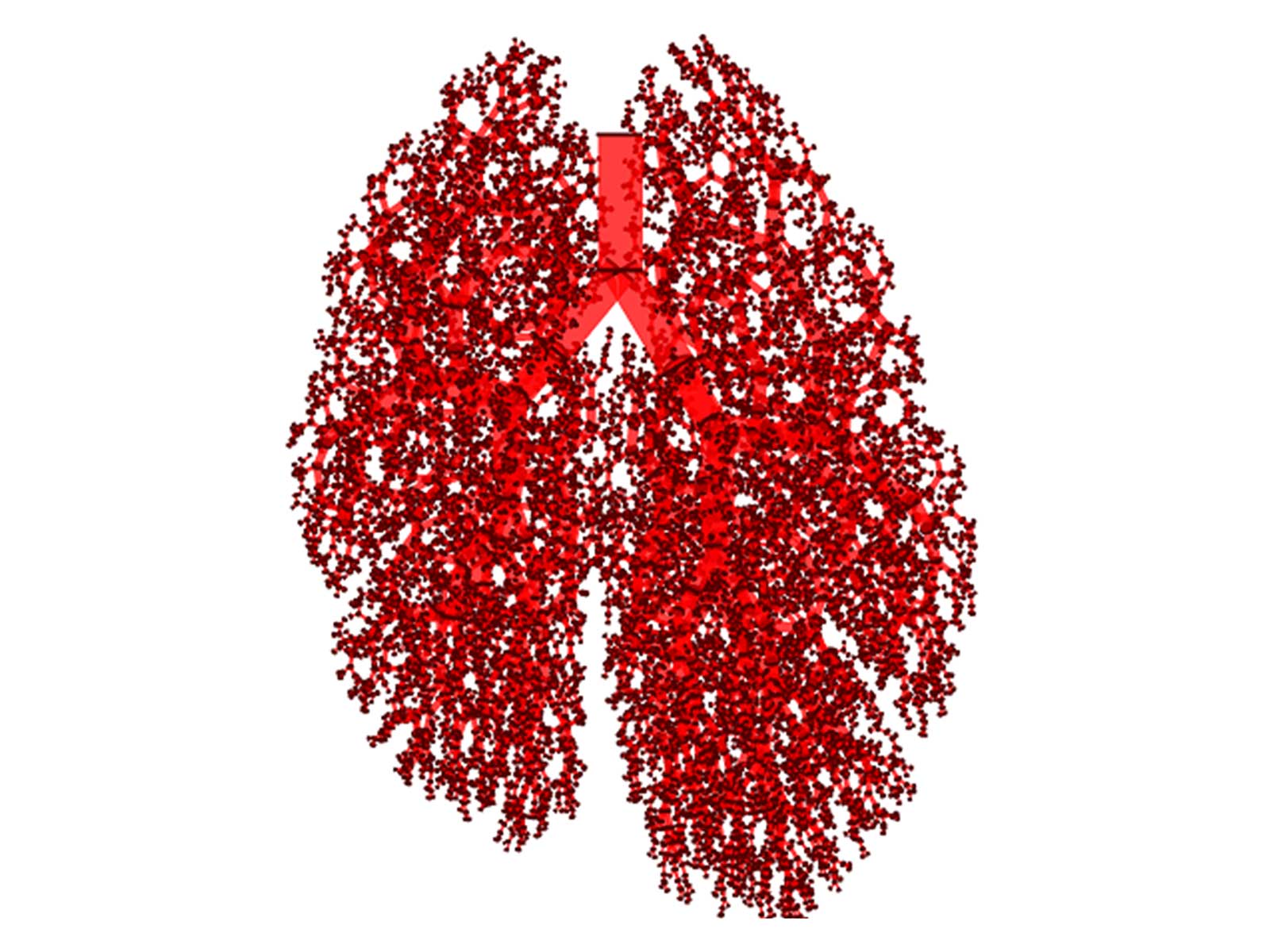
The researchers have started from the hypothesis that endogenous reinfection plays an important role in maintaining latent infection. "So, we have developed a model based on agents that describe the growth, fusion and proliferation of tuberculosis lesions in a computational bronchial tree, built from an iterative algorithm that generates the bronchial tube and bifurcations within a 3D volume of the lung surface area," explains Clara Prats, corresponding author of the article. The model has been provided with data and the parameters have been set using experimental data from computational tomography (CT) scans from five mini-pigs in the initial stages of infection. From these images, the lung surface area of the mini-pigs has been obtained in order to generate the computational lung. The size and placement of each of the TB lesions has also been obtained, which has allowed the researchers to adjust the model based on agents. "The result is a model that allows us to reproduce and understand the experimental data on the computer. We have been able to show an important relationship between the final number of lesions from tuberculosis with the frequency of endogenous reinfection and the growth of the lesions," adds Martí Català, the first author.
The model was also used as an in silico platform to explore the transition from latent infection to active disease, identifying the principal triggering factors: an elevated inflammatory reaction and the combination of a moderate inflammatory response with low respiratory amplitude. "This research is important because the structure of the lung of the minipig is very similar to that in humans, so that this model will allow us to make predictions for future interventions such as new biomarkers, preventive strategies and therapies for tuberculosis in humans," comments Pere-Joan Cardona, co-author of the project.
In silico computational models
The use of animal models, known as comparative medicine, has been one of the principal keys in the huge developments in medicine in the 20th century and particularly in the fields of life sciences and health for developing new treatments and drugs. New technologies have allowed for the refinement of this practice and this is main objective of the facilities, research and projects at the Centre for Comparative Medicine and Bioimage (CMCiB). The centre promotes research based on the Principle of the 3Rs: replace, reduce and refine. The use of computational tools for simulations, such as in this case, is a clear example. "Less is more in comparative medicine. Our group has been using alternative models for many years in pre-clinical research, such as the Drosophila model for tuberculosis research, a paradigmatic example. Now the possibilities offered by generated by computer are enormous," explains Cardona, Principal Researcher in the Experimental Tuberculosis Unit (UTE) and Scientific Director of the CMCiB. "In this case, we have obtained reliable results generated from a minimal number of animals and bioimaging tools. Now we will not need more animals, the model is in the computer and will serve us for a many future studies," he adds.
These are the first studies published within the framework of the 3Rs Programme at the CMCiB, which is supported by the "la Caixa" Foundation. Other projects are also underway on tuberculosis, ictus, computational models for HIV vaccines and to find new therapeutic targets in acute leukaemia.
Original article
Modelling the dynamics of tuberculosis lesions in a virtual lung: Role of the bronchial tree in endogenous reinfection.
Martí Català, Jordi Bechini, Montserrat Tenesa, Ricardo Pérez, Mariano Moya, Cristina Vilaplana, Joaquim Valls, Sergio Alonso, Daniel López, Pere-Joan Cardona, Clara Prats
PLOS Computational Biology,Published: May 20, 2020, doi.org/10.1371/journal.pcbi.1007772

